Home

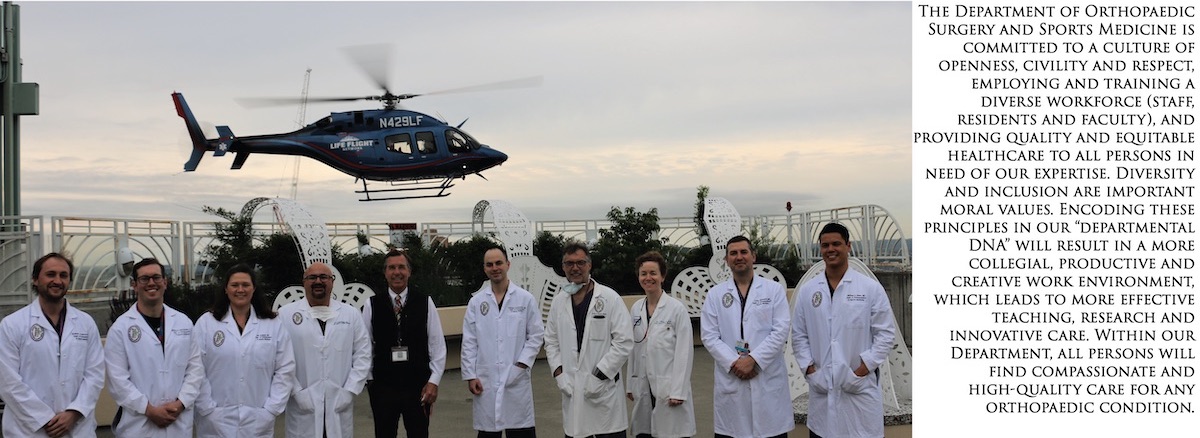
The University of Washington Department of Orthopaedic Surgery and Sports Medicine is committed to a culture of openness, civility and respect, employing and training a diverse workforce (staff, residents and faculty), and providing quality and equitable healthcare to all persons in need of our expertise.

The University of Washington, Department of Orthopaedic Surgery and Sports Medicine is committed to improving diversity not only in our department but in our orthopedics community as a whole. Learn more at the links below about our efforts to improve diversity and inclusivity in our department and become leaders at UW Medicine and in the orthopaedics community to better reach under-represented students.